File:Demethylation of 5-methylcytosine.svg
From Wikimedia Commons, the free media repository
Jump to navigation
Jump to search

Size of this PNG preview of this SVG file: 463 × 599 pixels. Other resolutions: 185 × 240 pixels | 371 × 480 pixels | 593 × 768 pixels | 791 × 1,024 pixels | 1,582 × 2,048 pixels | 816 × 1,056 pixels.
Original file (SVG file, nominally 816 × 1,056 pixels, file size: 127 KB)
File information
Structured data
Captions
Captions
Demethylation pathways for 5-Methylcytosine
Summary[edit]
| DescriptionDemethylation of 5-methylcytosine.svg |
English: Demethylation of 5-Methylcytosine (5mC) in DNA. As reviewed in 2018,[1] 5mC is oxidized by the ten-eleven translocation (TET) family of dioxygenases (TET1, TET2, TET3) to generate 5-hydroxymethylcytosine (5hmC). In successive steps TET enzymes further hydroxylate 5hmC to generate 5-formylcytosine (5fC) and 5-carboxylcytosine (5caC). Thymine-DNA glycosylase (TDG) recognizes the intermediate bases 5fC and 5caC and excises the glycosidic bond resulting in an apyrimidinic site (AP site). In an alternative oxidative deamination pathway, 5hmC can be oxidatively deaminated by activity-induced cytidine deaminase/apolipoprotein B mRNA editing complex (AID/APOBEC) deaminases to form 5-hydroxymethyluracil (5hmU) or 5mC can be converted to thymine (Thy). 5hmU can be cleaved by TDG, single-strand-selective monofunctional uracil-DNA glycosylase 1 (SMUG1), Nei-Like DNA Glycosylase 1 (NEIL1), or methyl-CpG binding protein 4 (MBD4). AP sites and T:G mismatches are then repaired by base excision repair (BER) enzymes to yield cytosine (Cyt). |
| Date | |
| Source | Own work |
| Author | Bernstein0275, User:Innerstream |
Licensing[edit]
I, the copyright holder of this work, hereby publish it under the following license:
This file is licensed under the Creative Commons Attribution-Share Alike 4.0 International license.
- You are free:
- to share – to copy, distribute and transmit the work
- to remix – to adapt the work
- Under the following conditions:
- attribution – You must give appropriate credit, provide a link to the license, and indicate if changes were made. You may do so in any reasonable manner, but not in any way that suggests the licensor endorses you or your use.
- share alike – If you remix, transform, or build upon the material, you must distribute your contributions under the same or compatible license as the original.
- ↑ (2018). "The Role of Activity-Dependent DNA Demethylation in the Adult Brain and in Neurological Disorders". Front Mol Neurosci 11: 169. DOI:10.3389/fnmol.2018.00169. PMID 29875631. PMC: 5975432.
File history
Click on a date/time to view the file as it appeared at that time.
| Date/Time | Thumbnail | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 13:01, 30 October 2021 |  | 816 × 1,056 (127 KB) | Innerstream (talk | contribs) | corrected and upgraded chemical structures |
| 17:32, 3 September 2018 | 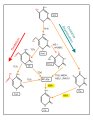 | 816 × 1,056 (41 KB) | Bernstein0275 (talk | contribs) | User created page with UploadWizard |
You cannot overwrite this file.
File usage on Commons
The following page uses this file:
File usage on other wikis
The following other wikis use this file:
- Usage on el.wikipedia.org
- Usage on en.wikipedia.org
- Usage on gl.wikipedia.org
- Usage on nl.wikipedia.org
- Usage on pt.wikipedia.org
- Usage on uk.wikipedia.org
Metadata
This file contains additional information such as Exif metadata which may have been added by the digital camera, scanner, or software program used to create or digitize it. If the file has been modified from its original state, some details such as the timestamp may not fully reflect those of the original file. The timestamp is only as accurate as the clock in the camera, and it may be completely wrong.
| Width | 816 |
|---|---|
| Height | 1056 |
Structured data
Items getoond in dit bestand
depicts
Waarde zonder Wikidata-item
3 September 2018
image/svg+xml
Hidden categories: