Category:Epigenetics
Przejdź do nawigacji
Przejdź do wyszukiwania
| Category Epigenetics on sister projects: | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
en:
|
||||||||||||||||||||||||
Podkategorie
Poniżej wyświetlono 9 spośród wszystkich 9 podkategorii tej kategorii.
D
- DNA methylation (64 pliki)
M
- Media from Epigenetics & Chromatin (13 plików)
S
- Epigenetics of schizophrenia (10 plików)
- Somaclonal variation (1 plik)
T
- TET enzymes (20 plików)
V
- Vitrovariation (1 plik)
Pliki w kategorii „Epigenetics”
Poniżej wyświetlono 125 spośród wszystkich 125 plików w tej kategorii.
-
13059 2015 737 Fig1 HTML.gif 719 × 992; 87 KB
-
Aging Cell - 2024 - Koncevičius - Epigenetic age oscillates during the day.pdf 1239 × 1629, 8 stron; 2,34 MB
-
Agouti Mice.jpg 1200 × 941; 601 KB
-
Beck–Fahrner syndrome or TET3 deficiency (episignature).jpg 1796 × 1841; 422 KB
-
Beck–Fahrner syndrome or TET3 deficiency episignature.png 2052 × 1755; 846 KB
-
Brian in a workshop.png 791 × 545; 840 KB
-
Cancer genetics-epigenetics yin-yang.jpg 1800 × 1790; 163 KB
-
Chemical structures of TET-oxidized products.jpg 839 × 590; 129 KB
-
Craniofacial features in Beck–Fahrner syndrome.png 1221 × 1638; 2,62 MB
-
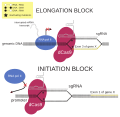
CRISPR Sterics.svg 1140 × 1105; 155 KB
-
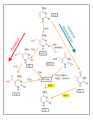
Crosstalk between metabolic and epigenetic regulation.jpg 2693 × 1277; 251 KB
-
Cytosine and 5-methylcytosine.jpg 1524 × 954; 91 KB
-
Cytosine and 5-methylcytosine.svg 610 × 373; 21 KB
-
Dam Methylase.png 1700 × 999; 466 KB
-
Demethylation of 5-methylcytosine.jpg 2550 × 3300; 1,36 MB
-
Demethylation of 5-methylcytosine.svg 816 × 1056; 127 KB
-
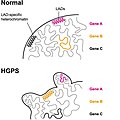
Deregulation LAD.jpg 1038 × 1042; 113 KB
-
Diagram Damage to Cancer Wiki 300dpi Workaround.svg 512 × 675; 314 KB
-
Diagram Damage to Cancer Wiki 300dpi(hy).png 683 × 900; 84 KB
-
Diagram Damage to Cancer Wiki 300dpi.svg 512 × 675; 4 KB
-
Differential-epigenetic-reprogramming-in-response-to-specific-endocrine-therapies-promotes-ncomms10044-s2.ogv 20 s, 1344 × 1024; 15,49 MB
-
Differential-epigenetic-reprogramming-in-response-to-specific-endocrine-therapies-promotes-ncomms10044-s3.ogv 20 s, 1344 × 1024; 12,31 MB
-
Differential-epigenetic-reprogramming-in-response-to-specific-endocrine-therapies-promotes-ncomms10044-s4.ogv 20 s, 1344 × 1024; 10,26 MB
-
Differential-epigenetic-reprogramming-in-response-to-specific-endocrine-therapies-promotes-ncomms10044-s5.ogv 19 s, 1344 × 1024; 18,75 MB
-
-
-
Differential-epigenetic-reprogramming-in-response-to-specific-endocrine-therapies-promotes-ncomms10044-s8.ogv 13 s, 1344 × 1024; 10,57 MB
-
-
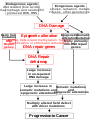
DNA damage, repair, alteration of repair in cancer.jpg 2550 × 3300; 2 MB
-
DNA damage, repair, epigenetic alteration of repair in cancer.jpg 2404 × 2628; 709 KB
-
DNA methylation (corrected).png 858 × 761; 37 KB
-
DNA methylation.jpg 6400 × 4800; 1,11 MB
-
Dosage compensation-ru.jpg 2291 × 1691; 2,04 MB
-
Dosage compensation.jpg 2291 × 1691; 661 KB
-
EffectorWiki.pdf 6900 × 6366; 393 KB
-
Epigen.png 800 × 515; 121 KB
-
EpigenByYear 1.png 3705 × 2272; 439 KB
-
Epigenetic alterations in tumour progression.svg 387 × 145; 347 KB
-
Epigenetic Inheritance Through The Female Line.png 504 × 400; 149 KB
-
Epigenetic Landscape - Waddington's Adaptation.png 944 × 1028; 468 KB
-
Epigenetic priming in Cancer.png 4096 × 1443; 989 KB
-
Epigenetic priming model.png 4096 × 4096; 1,75 MB
-
-
Epigenetic-Regulation-of-Histone-H3-Serine-10-Phosphorylation-Status-by-HCF-1-Proteins-in-C.-pone.0001213.s003.ogv 1 min 3 s, 348 × 319; 4,96 MB
-
Epigenetic-Regulation-of-Histone-H3-Serine-10-Phosphorylation-Status-by-HCF-1-Proteins-in-C.-pone.0001213.s004.ogv 1 min 3 s, 412 × 418; 5,95 MB
-
Epigenetic-Regulation-of-Histone-H3-Serine-10-Phosphorylation-Status-by-HCF-1-Proteins-in-C.-pone.0001213.s005.ogv 1 min 38 s, 344 × 206; 8,25 MB
-
Epigenetics modifications.png 574 × 324; 112 KB
-
Epigenetik Mekanizmalar.jpg 1068 × 800; 468 KB
-
Epigenetische Mechanismen.png 2187 × 1488; 2,04 MB
-
Epigenome-transparent-upscale.png 1203 × 1060; 562 KB
-
Epigenome.png 397 × 325; 72 KB
-
Epigentics 1.png 200 × 200; 5 KB
-
Epiphenotyping workflow1.png 2300 × 1170; 272 KB
-
Epiphenotyping.png 1836 × 1070; 221 KB
-
Examples of epiphenotyping applications.jpg 960 × 540; 75 KB
-
Fam40b-is-required-for-lineage-commitment-of-murine-embryonic-stem-cells-cddis2014273x4.ogv 21 s, 1024 × 768; 677 KB
-
Fam40b-is-required-for-lineage-commitment-of-murine-embryonic-stem-cells-cddis2014273x5.ogv 20 s, 1024 × 768; 344 KB
-
Gene regulation by TET proteins.png 1438 × 935; 491 KB
-
Genetic-epigenesis-pbio.1001325.s014.ogv 3,0 s, 1548 × 616; 254 KB
-
Genetic-epigenesis-pbio.1001325.s015.ogv 3,0 s, 608 × 302; 48 KB
-
Genetic-epigenesis-pbio.1001325.s016.ogv 4,0 s, 400 × 540; 69 KB
-
Genetic-epigenesis-pbio.1001325.s017.ogv 7,3 s, 1302 × 626; 187 KB
-
Genetic-epigenesis-pbio.1001325.s018.ogv 7,3 s, 1302 × 526; 143 KB
-
Genomic imprinting.svg 964 × 885; 552 KB
-
HDACs' role.jpg 1084 × 874; 82 KB
-
Heritable-Stochastic-Switching-Revealed-by-Single-Cell-Genealogy-pbio.0050239.sv001.ogv 15 s, 738 × 700; 526 KB
-
Heterochromatin-Formation-Promotes-Longevity-and-Represses-Ribosomal-RNA-Synthesis-pgen.1002473.s005.ogv 1 min 14 s, 352 × 288; 1,67 MB
-
Heterochromatin-Formation-Promotes-Longevity-and-Represses-Ribosomal-RNA-Synthesis-pgen.1002473.s006.ogv 1 min 10 s, 352 × 288; 1,7 MB
-
Heterochromatin-Formation-Promotes-Longevity-and-Represses-Ribosomal-RNA-Synthesis-pgen.1002473.s007.ogv 1 min 11 s, 352 × 288; 1,51 MB
-
Histone acetylation in regulating glial response after CNS injury.jpg 1026 × 782; 580 KB
-
Igf2 imprinting.svg 528 × 340; 60 KB
-
Ijms-20-03123-g001-550.webp 550 × 455; 35 KB
-
Inactivacio-cromosoma-x.svg 900 × 700; 61 KB
-
Initiation of DNA demethylation at a CpG site.svg 643 × 739; 38 KB
-
Intergenerational inheritance.png 479 × 338; 24 KB
-
Isoformes del DNMT3B.jpg 960 × 540; 57 KB
-
KDM5B.jpg 769 × 479; 133 KB
-
Kit paramutation.tif 1994 × 3699; 1,61 MB
-
Mechanismen epigenetischer Genregulation.webp 2055 × 1254; 114 KB
-
Methylation levels during mouse early embryonic development.jpg 816 × 1056; 59 KB
-
Methylation levels during mouse very early embryonic development.jpg 651 × 315; 37 KB
-
Model działania lunazyny na epigenom.png 689 × 572; 47 KB
-
Model działania lunazyny na epigenom2.png 689 × 572; 46 KB
-
-
-
-
-
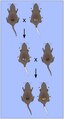
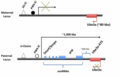
Non-Mendelian inheritance of mouse paramutations.png 1994 × 3699; 890 KB
-
Normal-cancer-epigenome.png 1007 × 1123; 185 KB
-
Normal-cancer-epigenome.svg 403 × 449; 502 KB
-
NucleosomeRegulation.png 1600 × 827; 235 KB
-
Parthenogenesis - Bischoff, Steve R 2010.tif 3000 × 1941; 39,49 MB
-
PD-1, PD-L1 and CTLA-4 targets of immune checkpoint inhibitors.png 1244 × 922; 602 KB
-
Peto-cromosomes-X.svg 744 × 1052; 72 KB
-
PKD-1 epigenetics regulation.jpg 800 × 489; 76 KB
-
Regulation of lymphoid development and function by TET proteins in the mouse.png 1126 × 1359; 523 KB
-
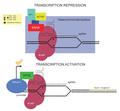
Regulation of transcription in mammals.jpg 2524 × 1372; 227 KB
-
Sc omics summary.svg 704 × 581; 104 KB
-
Schematic diagram of epigenetic mechanisms.jpg 850 × 745; 95 KB
-
Segragation.svg 981 × 607; 8 KB
-
Shelley Berger for the National Institutes of Health.jpg 867 × 849; 107 KB
-
Somaclonal variation and epigenetics.pdf 1239 × 1754, 116 stron; 6,64 MB
-
STARR-seq — Principles.png 1367 × 565; 206 KB
-
Targets of the epigenetic drugs.png 1370 × 641; 481 KB
-
TET enzymes and 5hmC in epigenetic regulation of mammalian neurobiology.jpg 3730 × 1694; 447 KB
-
TET enzymes in the process of DNA demethylation.jpg 1097 × 646; 125 KB
-
TET enzymes target signalling pathways.jpg 807 × 901; 257 KB
-
TET-mediated DNA modifications and demethylation.png 1587 × 1115; 358 KB
-
Transgenerational inheritance.png 629 × 315; 27 KB
-
Vorinostat Combined with DNMTi Epigenetically Controls the Proliferation of Lung Cancer Cells A549.pdf 1239 × 1752, 6 stron; 728 KB
-
W Reik photo cropped.png 568 × 862; 541 KB
-
Work flow for epigenome wide association study..png 685 × 600; 46 KB
-
X chromosome inactivation v3.png 1031 × 1057; 184 KB
-
Ген KDM6B и Церебральная Фолатная Недостаточность - Han et al 2023.pdf 1239 × 1752, 17 stron; 1,69 MB