Category:Protein folding
Vai alla navigazione
Vai alla ricerca
the process of assisting in the covalent and noncovalent assembly of single chain polypeptides or multisubunit complexes into the correct tertiary structure | |||||
| Carica un file multimediale | |||||
| Istanza di | |||||
|---|---|---|---|---|---|
| Sottoclasse di | |||||
| |||||
Sottocategorie
Questa categoria contiene le 7 sottocategorie indicate di seguito, su un totale di 7.
File nella categoria "Protein folding"
Questa categoria contiene 200 file, indicati di seguito, su un totale di 252.
(pagina precedente) (pagina successiva)-
1X9J.png 640 × 434; 364 KB
-
3bq.ent.pdb.png 385 × 388; 59 KB
-
41594 2023 1201 Fig4.webp 1 826 × 2 181; 513 KB
-
41594 2023 1201 FigE10 Fig15.webp 2 100 × 737; 117 KB
-
41594 2023 1201 FigE2 Fig7.webp 2 086 × 2 268; 217 KB
-
41594 2023 1201 FigE3 Fig8.webp 1 805 × 1 989; 207 KB
-
41594 2023 1201 FigE4 Fig9.webp 2 105 × 1 886; 417 KB
-
41594 2023 1201 FigE5 Fig10.webp 1 858 × 2 712; 429 KB
-
41594 2023 1201 FigE6 Fig11.webp 2 072 × 2 189; 364 KB
-
41594 2023 1201 FigE7 Fig12.webp 2 079 × 1 678; 325 KB
-
41594 2023 1201 FigE8 Fig13.webp 1 651 × 2 031; 378 KB
-
41594 2023 1201 FigE9 Fig14.webp 2 086 × 1 531; 287 KB
-
4oq9 chainA jellyroll.png 1 024 × 1 024; 271 KB
-
A model of DBH active site based on PHM.jpg 360 × 670; 114 KB
-
A replica exchange molecular dynamics simulation of the folding of.webm 1 min 40 s, 3 872 × 2 142; 47,25 MB
-
-
-
-
-
A-Customized-Light-Sheet-Microscope-to-Measure-Spatio-Temporal-Protein-Dynamics-in-Small-Model-pone.0127869.s010.ogv 43 s, 1 110 × 1 000; 1,21 MB
-
-
A-SEL1L-Mutation-Links-a-Canine-Progressive-Early-Onset-Cerebellar-Ataxia-to-the-Endoplasmic-pgen.1002759.s007.ogv 1 min 37 s, 640 × 480; 19,73 MB
-
A-SEL1L-Mutation-Links-a-Canine-Progressive-Early-Onset-Cerebellar-Ataxia-to-the-Endoplasmic-pgen.1002759.s008.ogv 1 min 56 s, 640 × 480; 23,41 MB
-
A-SEL1L-Mutation-Links-a-Canine-Progressive-Early-Onset-Cerebellar-Ataxia-to-the-Endoplasmic-pgen.1002759.s009.ogv 1 min 48 s, 640 × 480; 18,26 MB
-
AAAA.jpg 240 × 240; 21 KB
-
AAAA1.jpg 240 × 240; 20 KB
-
AaHIT1 3D residue19-88.JPG 261 × 279; 21 KB
-
ACBP MSM from Folding@home.tiff 3 036 × 1 817; 1,1 MB
-
Alpha helix neg60 neg45 topview.png 572 × 566; 96 KB
-
Alpha sheet bonding schematic-color.svg 2 138 × 2 181; 50 KB
-
Alpha strand 50 50 vertical.png 287 × 820; 63 KB
-
Alpha strand 50 50.png 820 × 287; 64 KB
-
-
-
-
An-Automatic-Refolding-Apparatus-for-Preparative-Scale-Protein-Production-pone.0045891.s008.ogv 57 s, 1 920 × 1 080; 570 KB
-
An-Automatic-Refolding-Apparatus-for-Preparative-Scale-Protein-Production-pone.0045891.s009.ogv 54 s, 1 920 × 1 080; 540 KB
-
An-Automatic-Refolding-Apparatus-for-Preparative-Scale-Protein-Production-pone.0045891.s010.ogv 54 s, 1 920 × 1 080; 530 KB
-
An-Automatic-Refolding-Apparatus-for-Preparative-Scale-Protein-Production-pone.0045891.s011.ogv 54 s, 1 920 × 1 080; 522 KB
-
An-Automatic-Refolding-Apparatus-for-Preparative-Scale-Protein-Production-pone.0045891.s012.ogv 36 s, 1 920 × 1 080; 364 KB
-
An-Automatic-Refolding-Apparatus-for-Preparative-Scale-Protein-Production-pone.0045891.s013.ogv 54 s, 1 920 × 1 080; 545 KB
-
An-Automatic-Refolding-Apparatus-for-Preparative-Scale-Protein-Production-pone.0045891.s014.ogv 21 s, 1 920 × 1 080; 256 KB
-
An-Automatic-Refolding-Apparatus-for-Preparative-Scale-Protein-Production-pone.0045891.s015.ogv 39 s, 1 920 × 1 080; 400 KB
-
-
-
An-Essential-Nonredundant-Role-for-Mycobacterial-DnaK-in-Native-Protein-Folding-pgen.1004516.s019.ogv 10 s, 1 050 × 350; 177 KB
-
-
-
Aquagliceroporina.png 604 × 338; 159 KB
-
Artificial-targeting-of-misfolded-cytosolic-proteins-to-endoplasmic-reticulum-as-a-mechanism-for-srep12088-s2.ogv 5,8 s, 1 024 × 1 024; 3,56 MB
-
Artificial-targeting-of-misfolded-cytosolic-proteins-to-endoplasmic-reticulum-as-a-mechanism-for-srep12088-s3.ogv 8,1 s, 1 024 × 1 024; 1,59 MB
-
Artificial-targeting-of-misfolded-cytosolic-proteins-to-endoplasmic-reticulum-as-a-mechanism-for-srep12088-s4.ogv 9,6 s, 1 024 × 1 024; 1,14 MB
-
-
-
BCLFold---De-Novo-Prediction-of-Complex-and-Large-Protein-Topologies-by-Assembly-of-Secondary-pone.0049240.s011.ogv 1 min 2 s, 720 × 480; 5,63 MB
-
BCLFold---De-Novo-Prediction-of-Complex-and-Large-Protein-Topologies-by-Assembly-of-Secondary-pone.0049240.s012.ogv 2 min 22 s, 720 × 480; 5,1 MB
-
BCLFold---De-Novo-Prediction-of-Complex-and-Large-Protein-Topologies-by-Assembly-of-Secondary-pone.0049240.s013.ogv 6 min 12 s, 720 × 480; 8,99 MB
-
BCLFold---De-Novo-Prediction-of-Complex-and-Large-Protein-Topologies-by-Assembly-of-Secondary-pone.0049240.s014.ogv 5 min 43 s, 720 × 480; 6,77 MB
-
BCLFold---De-Novo-Prediction-of-Complex-and-Large-Protein-Topologies-by-Assembly-of-Secondary-pone.0049240.s015.ogv 1 min 17 s, 720 × 480; 7,63 MB
-
-
BCLFold---De-Novo-Prediction-of-Complex-and-Large-Protein-Topologies-by-Assembly-of-Secondary-pone.0049240.s017.ogv 2 min 51 s, 720 × 480; 5,91 MB
-
BCLFold---De-Novo-Prediction-of-Complex-and-Large-Protein-Topologies-by-Assembly-of-Secondary-pone.0049240.s018.ogv 6 min 22 s, 720 × 480; 8,99 MB
-
BCLFold---De-Novo-Prediction-of-Complex-and-Large-Protein-Topologies-by-Assembly-of-Secondary-pone.0049240.s019.ogv 4 min 50 s, 720 × 480; 5,61 MB
-
-
-
BCLFold---De-Novo-Prediction-of-Complex-and-Large-Protein-Topologies-by-Assembly-of-Secondary-pone.0049240.s022.ogv 2 min 52 s, 720 × 480; 4,65 MB
-
BCLFold---De-Novo-Prediction-of-Complex-and-Large-Protein-Topologies-by-Assembly-of-Secondary-pone.0049240.s023.ogv 8 min 13 s, 720 × 480; 9,05 MB
-
BCLFold---De-Novo-Prediction-of-Complex-and-Large-Protein-Topologies-by-Assembly-of-Secondary-pone.0049240.s024.ogv 5 min 5 s, 720 × 480; 4,79 MB
-
Beta sheet bonding antiparallel-color.svg 1 151 × 3 855; 15 KB
-
Beta sheet bonding parallel-color.svg 1 151 × 3 855; 49 KB
-
Beta-meander1.png 393 × 255; 24 KB
-
BetaPleatedSheetProtein.png 610 × 422; 38 KB
-
-
-
-
-
Bj1.png 902 × 536; 371 KB
-
Bj2.png 452 × 611; 110 KB
-
Bj3.png 412 × 544; 69 KB
-
BPTI seq ribbon sticks.jpg 1 024 × 768; 246 KB
-
C-FLIPl.png 1 781 × 1 283; 306 KB
-
CalmodulinConformation.png 700 × 671; 149 KB
-
Chaperonin 1AON.png 1 956 × 850; 1,55 MB
-
Circular-solenoid-domain-sketch.png 662 × 582; 53 KB
-
-
Comparison of ferritin and transferrin.png 1 148 × 580; 436 KB
-
-
Correct-folding-of-an-α-helix-and-a-β-hairpin-using-a-polarized-2D-torsional-potential-srep10359-s2.ogv 12 min 0 s, 384 × 288; 17,92 MB
-
Coupled-Protein-Diffusion-and-Folding-in-the-Cell-pone.0113040.s002.ogv 4,0 s, 103 × 124; 18 KB
-
De-Novo-Generation-of-Infectious-Prions-In-Vitro-Produces-a-New-Disease-Phenotype-ppat.1000421.s004.ogv 3 min 2 s, 320 × 240; 14,98 MB
-
De-Novo-Generation-of-Infectious-Prions-In-Vitro-Produces-a-New-Disease-Phenotype-ppat.1000421.s005.ogv 3 min 19 s, 320 × 240; 17,24 MB
-
-
-
-
Development-of-Cysteine-Free-Fluorescent-Proteins-for-the-Oxidative-Environment-pone.0037551.s004.ogv 8,9 s, 512 × 512; 3,85 MB
-
Diacylglycerol lipase α (via AlphaFold2, ColabFold, and PyMol).png 3 016 × 1 566; 715 KB
-
Diacylglycerol lipase β (via AlphaFold2, ColabFold, and PyMol).png 3 016 × 1 566; 695 KB
-
-
-
Dimer of trimer 1YR3.png 460 × 400; 141 KB
-
DisulfideBondFormation.png 786 × 242; 3 KB
-
DisulfideBondFormation.svg 957 × 407; 50 KB
-
Divergent-Effects-of-PERK-and-IRE1-Signaling-on-Cell-Viability-pone.0004170.s004.ogv 39 s, 696 × 260; 12,69 MB
-
Divergent-Effects-of-PERK-and-IRE1-Signaling-on-Cell-Viability-pone.0004170.s005.ogv 40 s, 696 × 260; 9,61 MB
-
Divergent-Effects-of-PERK-and-IRE1-Signaling-on-Cell-Viability-pone.0004170.s006.ogv 40 s, 696 × 260; 19,67 MB
-
DsRBD.png 596 × 638; 255 KB
-
-
-
Dysferlin-Peptides-Reallocate-Mutated-Dysferlin-Thereby-Restoring-Function-pone.0049603.s007.ogv 5,0 s, 512 × 512; 117 KB
-
Dysferlin-Peptides-Reallocate-Mutated-Dysferlin-Thereby-Restoring-Function-pone.0049603.s008.ogv 5,0 s, 512 × 512; 28 KB
-
Dysferlin-Peptides-Reallocate-Mutated-Dysferlin-Thereby-Restoring-Function-pone.0049603.s009.ogv 5,0 s, 512 × 512; 16 KB
-
Dysferlin-Peptides-Reallocate-Mutated-Dysferlin-Thereby-Restoring-Function-pone.0049603.s010.ogv 5,0 s, 512 × 504; 77 KB
-
Dysferlin-Peptides-Reallocate-Mutated-Dysferlin-Thereby-Restoring-Function-pone.0049603.s011.ogv 56 s, 800 × 602; 2,54 MB
-
Dysferlin-Peptides-Reallocate-Mutated-Dysferlin-Thereby-Restoring-Function-pone.0049603.s012.ogv 1 min 16 s, 800 × 602; 3,22 MB
-
Dysferlin-Peptides-Reallocate-Mutated-Dysferlin-Thereby-Restoring-Function-pone.0049603.s013.ogv 25 s, 800 × 602; 1,55 MB
-
Dysferlin-Peptides-Reallocate-Mutated-Dysferlin-Thereby-Restoring-Function-pone.0049603.s014.ogv 52 s, 896 × 672; 2,63 MB
-
Estructura proteínas.png 474 × 536; 26 KB
-
Evolutionarily-Conserved-Linkage-between-Enzyme-Fold-Flexibility-and-Catalysis-pbio.1001193.s013.ogv 1,4 s, 1 280 × 960; 1,37 MB
-
Evolutionarily-Conserved-Linkage-between-Enzyme-Fold-Flexibility-and-Catalysis-pbio.1001193.s014.ogv 1,4 s, 1 280 × 960; 1,26 MB
-
Evolutionarily-Conserved-Linkage-between-Enzyme-Fold-Flexibility-and-Catalysis-pbio.1001193.s015.ogv 1,4 s, 1 280 × 960; 1,27 MB
-
Evolutionarily-Conserved-Linkage-between-Enzyme-Fold-Flexibility-and-Catalysis-pbio.1001193.s016.ogv 1,4 s, 1 024 × 768; 1,18 MB
-
Evolutionarily-Conserved-Linkage-between-Enzyme-Fold-Flexibility-and-Catalysis-pbio.1001193.s017.ogv 1,4 s, 1 024 × 768; 1,37 MB
-
Evolutionarily-Conserved-Linkage-between-Enzyme-Fold-Flexibility-and-Catalysis-pbio.1001193.s018.ogv 1,4 s, 1 024 × 768; 1,23 MB
-
Evolutionarily-Conserved-Linkage-between-Enzyme-Fold-Flexibility-and-Catalysis-pbio.1001193.s019.ogv 1,4 s, 1 280 × 960; 1,23 MB
-
Evolutionarily-Conserved-Linkage-between-Enzyme-Fold-Flexibility-and-Catalysis-pbio.1001193.s020.ogv 1,4 s, 1 280 × 960; 1,24 MB
-
Evolutionarily-Conserved-Linkage-between-Enzyme-Fold-Flexibility-and-Catalysis-pbio.1001193.s021.ogv 1,4 s, 1 280 × 960; 1,25 MB
-
Experimental-and-Computational-Analysis-of-Polyglutamine-Mediated-Cytotoxicity-pcbi.1000944.s004.ogv 14 s, 694 × 520; 3,95 MB
-
-
Experimental-and-Computational-Analysis-of-Polyglutamine-Mediated-Cytotoxicity-pcbi.1000944.s006.ogv 14 s, 694 × 520; 5,08 MB
-
-
Experimental-and-Computational-Analysis-of-Polyglutamine-Mediated-Cytotoxicity-pcbi.1000944.s008.ogv 14 s, 694 × 520; 5,87 MB
-
-
Experimental-and-Computational-Analysis-of-Polyglutamine-Mediated-Cytotoxicity-pcbi.1000944.s010.ogv 14 s, 694 × 520; 4,67 MB
-
-
Exploring-the-Universe-of-Protein-Structures-beyond-the-Protein-Data-Bank-pcbi.1000957.s008.ogv 0,0 s, 656 × 832; 6,59 MB
-
-
-
-
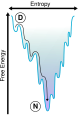
Folding energy diagram terbidium4.jpg 507 × 583; 29 KB
-
Folding Funnel.svg 167 × 255; 10 KB
-
-
FRET-Assisted-Determination-of-CLN3-Membrane-Topology-pone.0102593.s001.ogv 25 s, 1 536 × 512; 5,26 MB
-
Hexosaminidase A (heterodimer, with van der Waals interactions).jpg 700 × 400; 220 KB
-
HLA-A*02 protein structure.jpg 500 × 500; 44 KB
-
-
-
-
-
HP Lattice Protein Model Schematic.png 720 × 540; 49 KB
-
I-TASSER predicted protein fold of Fam188a.gif 535 × 535; 25 KB
-
Isomerization pathway in bacteria (mechanism).jpg 599 × 414; 20 KB
-
Isomerization pathway in bacteria(mechanism).jpg 541 × 372; 17 KB
-
KSI homodimer.png 640 × 434; 119 KB
-
Linear-solenoid-domain-sketch.png 662 × 582; 55 KB
-
Magnetotactic-molecular-architectures-from-self-assembly-of-β-peptide-foldamers-ncomms9747-s2.ogv 15 s, 640 × 480; 1,68 MB
-
-
Magnetotactic-molecular-architectures-from-self-assembly-of-β-peptide-foldamers-ncomms9747-s4.ogv 30 s, 640 × 480; 5,68 MB
-
Main protein structure levels cs.svg 463 × 808; 252 KB
-
-
MDC1-tandem-BRCT-domains-with-ligand.png 766 × 688; 104 KB
-
Membrane-Permeabilization-by-Oligomeric-α-Synuclein-In-Search-of-the-Mechanism-pone.0014292.s001.ogv 8,6 s, 512 × 512; 1,33 MB
-
-
-
-
-
Native-structure-based-modeling-and-simulation-of-biomolecular-systems-per-mouse-click-12859 2014 6561 MOESM1 ESM.ogv 1 min 56 s, 1 024 × 1 024; 7,32 MB
-
Native-structure-based-modeling-and-simulation-of-biomolecular-systems-per-mouse-click-12859 2014 6561 MOESM2 ESM.ogv 2 min 10 s, 1 024 × 1 024; 8,92 MB
-
Native-structure-based-modeling-and-simulation-of-biomolecular-systems-per-mouse-click-12859 2014 6561 MOESM3 ESM.ogv 1 min 36 s, 1 024 × 1 024; 7,94 MB
-
-
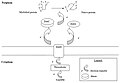
Oxidative folding in eukaryotes (mechanism).jpg 538 × 441; 20 KB
-
Oxidative folding in eukaryotes(mechanism).jpg 554 × 425; 20 KB
-
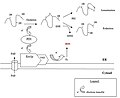
Oxidative pathway in bacteria (mechanism).jpg 619 × 448; 22 KB
-
Oxidative pathway in bacteria(mechanism).jpg 548 × 381; 19 KB
-
P3 peptide 2013-10-23 17-14.jpg 504 × 504; 56 KB
-
-
Protein fold.png 373 × 266; 47 KB
-
Protein Folding 2.JPG 370 × 260; 9 KB
-
Protein folding figure-es.png 1 042 × 1 232; 204 KB
-
Protein folding figure.png 1 042 × 1 232; 271 KB
-
Protein folding schematic.png 930 × 826; 43 KB
-
Protein folding.png 994 × 440; 70 KB
-
Protein Structural changes timescale matched with NMR experiments.png 1 416 × 544; 93 KB
-
Protein-structure hy.png 396 × 540; 64 KB
-
Protein-structure uk.jpg 287 × 515; 42 KB
-
Protein-structure.png 396 × 540; 49 KB
-
ProteinStructures fr.png 566 × 1 514; 116 KB
-
ProteinStructures.gif 396 × 540; 20 KB
-
ProteinStructures.png 566 × 1 700; 161 KB
-
RACK1.png 750 × 675; 220 KB
-
Ribbon diagram representation of the folding of the protein barnase.png 570 × 859; 127 KB
-
Ribose-5-phosphate isomerase.png 662 × 333; 223 KB
-
-
-
-
-
-
-
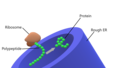
Rough ER Close up.png 1 188 × 672; 138 KB
-
SBPase crystallographic structure.png 647 × 555; 229 KB