Category:Adenosine triphosphatases
Bước tới điều hướng
Bước tới tìm kiếm
class of enzymes | |||||
| Tải lên phương tiện | |||||
| Là một |
| ||||
|---|---|---|---|---|---|
| Là tập hợp con của |
| ||||
| |||||
Thể loại con
Thể loại này có 4 thể loại con sau, trên tổng số 4 thể loại con.
S
V
Tập tin trong thể loại “Adenosine triphosphatases”
79 tập tin sau nằm trong thể loại này, trong tổng số 79 tập tin.
-
Adenylyl.jpg 355×433; 36 kB
-
An-Essential-Role-for-Katanin-p80-and-Microtubule-Severing-in-Male-Gamete-Production-pgen.1002698.s006.ogv 6,5 s, 1.024×1.024; 3,56 MB
-
An-Essential-Role-for-Katanin-p80-and-Microtubule-Severing-in-Male-Gamete-Production-pgen.1002698.s007.ogv 4,5 s, 1.024×1.024; 1,52 MB
-
An-Essential-Role-for-Katanin-p80-and-Microtubule-Severing-in-Male-Gamete-Production-pgen.1002698.s008.ogv 5,0 s, 1.024×1.024; 106 kB
-
-
Atp i la rotenona.jpg 680×324; 91 kB
-
ATPase cycle.png 612×526; 113 kB
-
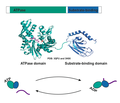
BiP’s Structure and ATPase Cycle.png 679×545; 116 kB
-
Clamp-loader-ATPases-and-the-evolution-of-DNA-replication-machinery-1741-7007-10-34-S1.ogv 23 s, 640×480; 6,22 MB
-
-
Clamp-loader-ATPases-and-the-evolution-of-DNA-replication-machinery-1741-7007-10-34-S3.ogv 5,0 s, 640×480; 1,69 MB
-
-
-
-
-
-
-
-
-
-
DHC ADP.webm 29 s, 624×606; 8,9 MB
-
-
Estructura de la P4-ATPasa.png 2.107×1.594; 340 kB
-
Estructura P4-ATPasa.png 3.700×2.081; 380 kB
-
Estrutura ATP13A2.jpg 1.055×795; 109 kB
-
Flexibility-within-the-Rotor-and-Stators-of-the-Vacuolar-H+-ATPase-pone.0082207.s003.ogv 1,4 s, 300×375; 199 kB
-
Flexibility-within-the-Rotor-and-Stators-of-the-Vacuolar-H+-ATPase-pone.0082207.s004.ogv 1,4 s, 300×375; 196 kB
-
Flexibility-within-the-Rotor-and-Stators-of-the-Vacuolar-H+-ATPase-pone.0082207.s005.ogv 1,8 s, 300×397; 263 kB
-
Flexibility-within-the-Rotor-and-Stators-of-the-Vacuolar-H+-ATPase-pone.0082207.s006.ogv 7,3 s, 1.024×1.024; 2,54 MB
-
Flexibility-within-the-Rotor-and-Stators-of-the-Vacuolar-H+-ATPase-pone.0082207.s007.ogv 0,0 s, 1.024×1.024; 2,17 MB
-
Flexibility-within-the-Rotor-and-Stators-of-the-Vacuolar-H+-ATPase-pone.0082207.s008.ogv 0,0 s, 800×800; 783 kB
-
Flexibility-within-the-Rotor-and-Stators-of-the-Vacuolar-H+-ATPase-pone.0082207.s009.ogv 1,7 s, 800×800; 440 kB
-
Flexibility-within-the-Rotor-and-Stators-of-the-Vacuolar-H+-ATPase-pone.0082207.s010.ogv 4,2 s, 1.108×1.108; 3,45 MB
-
Flexibility-within-the-Rotor-and-Stators-of-the-Vacuolar-H+-ATPase-pone.0082207.s011.ogv 4,2 s, 1.082×1.108; 3,55 MB
-
Flexibility-within-the-Rotor-and-Stators-of-the-Vacuolar-H+-ATPase-pone.0082207.s012.ogv 1,7 s, 1.250×1.109; 1,21 MB
-
Fo subunit of ATPase C1. Picture Created in PyMol.png 657×404; 245 kB
-
Fusion-of-Protein-Aggregates-Facilitates-Asymmetric-Damage-Segregation-pbio.1001886.s005.ogv 8,7 s, 212×218; 280 kB
-
Fusion-of-Protein-Aggregates-Facilitates-Asymmetric-Damage-Segregation-pbio.1001886.s006.ogv 8,7 s, 224×242; 239 kB
-
Gastric hydrogen potassium ATPase mechanism.svg 600×400; 2 kB
-
Mecanismes tranport atpasa.png 991×340; 146 kB
-
Mecanismes tranport flipasa.png 991×340; 146 kB
-
Molecular-snapshots-of-the-Pex16-AAA+-complex-in-action-ncomms8331-s2.ogv 0,8 s, 833×830; 288 kB
-
Noise-Induced-Min-Phenotypes-in-E.-coli-pcbi.0020080.sv001.ogv 34 s, 320×226; 5,13 MB
-
Noise-Induced-Min-Phenotypes-in-E.-coli-pcbi.0020080.sv002.ogv 20 s, 320×226; 319 kB
-
Noise-Induced-Min-Phenotypes-in-E.-coli-pcbi.0020080.sv003.ogv 34 s, 320×192; 3,53 MB
-
Noise-Induced-Min-Phenotypes-in-E.-coli-pcbi.0020080.sv004.ogv 41 s, 320×190; 419 kB
-
Noise-Induced-Min-Phenotypes-in-E.-coli-pcbi.0020080.sv005.ogv 27 s, 240×240; 5,16 MB
-
Noise-Induced-Min-Phenotypes-in-E.-coli-pcbi.0020080.sv006.ogv 6,8 s, 240×240; 157 kB
-
Noise-Induced-Min-Phenotypes-in-E.-coli-pcbi.0020080.sv007.ogv 33 s, 320×90; 2,94 MB
-
Noise-Induced-Min-Phenotypes-in-E.-coli-pcbi.0020080.sv008.ogv 12 s, 320×108; 111 kB
-
Noise-Induced-Min-Phenotypes-in-E.-coli-pcbi.0020080.sv009.ogv 1,2 s, 320×110; 17 kB
-
-
-
-
-
Pathways-of-allosteric-regulation-in-Hsp70-chaperones-ncomms9308-s2.ogv 2,7 s, 1.434×1.253; 1,41 MB
-
Pathways-of-allosteric-regulation-in-Hsp70-chaperones-ncomms9308-s3.ogv 2,7 s, 1.533×839; 1,31 MB
-
Pathways-of-allosteric-regulation-in-Hsp70-chaperones-ncomms9308-s4.ogv 16 s, 1.364×1.376; 4,01 MB
-
-
-
-
-
-
-
-
The-Hexameric-Structures-of-Human-Heat-Shock-Protein-90-pone.0019961.s001.ogv 7,7 s, 600×600; 12,69 MB
-
-
-
-
-
-
-
TumRacGAP-functions-as-a-switch-activating-the-Pavkinesin-6-motor-ncomms11182-s2.ogv 3,3 s, 320×160; 20 kB
-
TumRacGAP-functions-as-a-switch-activating-the-Pavkinesin-6-motor-ncomms11182-s3.ogv 4,9 s, 320×180; 35 kB
-
TumRacGAP-functions-as-a-switch-activating-the-Pavkinesin-6-motor-ncomms11182-s4.ogv 3,1 s, 320×180; 18 kB
-
TumRacGAP-functions-as-a-switch-activating-the-Pavkinesin-6-motor-ncomms11182-s5.ogv 5,7 s, 360×180; 39 kB
-
TumRacGAP-functions-as-a-switch-activating-the-Pavkinesin-6-motor-ncomms11182-s6.ogv 4,4 s, 320×180; 23 kB
-
TumRacGAP-functions-as-a-switch-activating-the-Pavkinesin-6-motor-ncomms11182-s7.ogv 3,0 s, 320×180; 14 kB
-
TumRacGAP-functions-as-a-switch-activating-the-Pavkinesin-6-motor-ncomms11182-s8.ogv 4,3 s, 320×180; 18 kB